Course overview
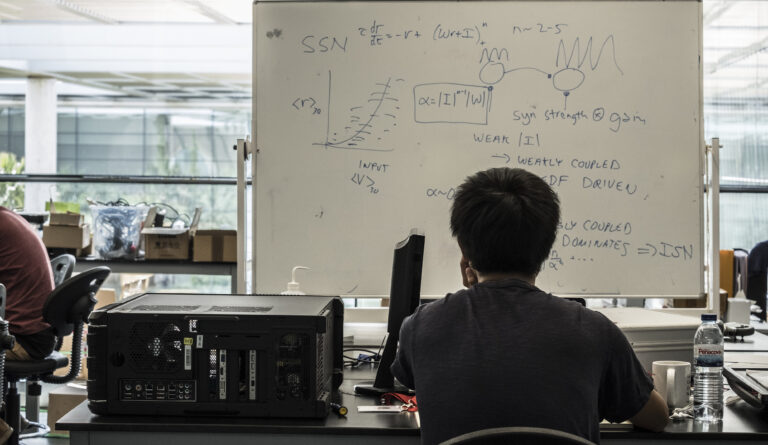
Understanding the links between activity in neural circuits and behavior is a fundamental problem in neuroscience. Attacking this problem requires detailed information about the cell types in neural circuits and their connectivity, and recording the spatiotemporal patterns of activity in the intact brain during behaviour. Furthermore, probing causal relationships between cellular and circuit-level processes and behaviour requires perturbation of specific elements of the circuit in a temporally and spatially precise manner.
This course will highlight the new anatomical, genetic, optical, electrophysiological, optogenetic, and pharmacogenetic approaches that are available for addressing these challenges. The faculty will discuss tool development through to their implementation in diverse model systems, including mice and zebrafish. Students will learn the potential and limitations of these techniques, allowing them to both design and interpret experiments correctly.
Course directors

Michael Hausser
Course Director
Wolfson Institute for Biomedical Research, UCL, UK

Claire Wyart
Course Director
Paris Brain Institute, ICM, France

Tiago Branco
Course Director
Sainsbury Wellcome Centre, UK

Susana Lima
Course Director
Champalimaud Foundation, Portugal
Keynote Speakers
Isaac Bianco, University College London, UK
Ed Boyden, Massachusetts Institute of Technology, USA
Michael Brecht, Bernstein Center for Computational Neuroscience, Germany
Megan Carey, Champalimaud Foundation, Portugal
Eugenia Chiappe, Champalimaud Foundation, Portugal
Winfried Denk, Max Planck Institute of Neurobiology, Germany
Emily Dennis, HHMI, Janelia, USA
Valentina Emiliani, Vision Institute, France
Ken Harris, University College London, UK
Greg Jefferis MRC Laboratory of Molecular Biology, UK
Na Ji, University of California, USA
Mackenzie Mathis, EPFL, Switzerland
Marta Moita, Champalimaud Foundation, Portugal
Francois St-Pierre, Baylor College of Medicine, USA
Michael Orger, Champalimaud Foundation, Portugal
Marius Pachitariu, HHMI, Janelia, USA
Darcy Peterka, Columbia University, USA
Pavan Ramdya, EPFL, Switzerland
Ana João Rodrigues, ICVS, Minho University, Portugal
Botond Roska, Institute of Molecular and Clinical Ophthalmology, Switzerland
Nick Steinmetz, University of Washington, USA
Carsen Stringer, HHMI, Janelia, USA
Karel Svoboda, HHMI, Janelia, USA
Scott Waddell, Oxford University, UK
Chris Xu, Cornell University, USA
Ofer Yizhar, Weizmann Institute of Science, Israel
Instructors
Dustin Herrmann, Wolfson Institute for Biomedical Research, UCL, UK
Maxime Beau, Wolfson Institute for Biomedical Research, UCL, UK
Edgar Baumler, Wolfson Institute for Biomedical Research, UCL, UK
Moritz Buchholz, Wolfson Institute for Biomedical Research, UCL, UK
Maha Dhanasekar, Paris Brain Institute, ICM, France
Pierre Tissier, Paris Brain Institute, ICM, France
Martin Carbó-Tano, Paris Brain Institute, ICM, France
Olivier Mirat, Paris Brain Institute, ICM, France
Dario Campagner, Sainsbury Wellcome Centre, UK
Tan YuLin, Sainsbury Wellcome Centre, UK
Hugo Marques, Champalimaud Foundation, Portugal
Ana Gonçalves, Champalimaud Foundation, Portugal
Course content
Schedule
Week 1
Sunday, June 19th: Arrival & Welcome reception
Monday, June 20th: Keynote lecture by Chris Xu / Ed Boyden Afternoon: Student presentations & Poster session I
Tuesday, June 21st: Keynote lecture by Eugenia Chiappe / Pavan Ramdya Afternoon: TA & Experimental Resources presentation & Poster session II
Wednesday, June 22nd: Keynote lecture by Greg Jefferis / Tiago Branco Afternoon TA & Experimental Resources presentation / Project Design & Discussion
Thursday, June 23rd: Morning tutorial by Scott Waddell / Na Ji Afternoon: Rig rotation / Project Design & Discussion
Friday, June 24th: Keynote lecture by Winfried Denk / Susana Lima Afternoon: Rig rotation/ Project Design & Discussion
Friday, June 25th: Surf/Beach Day
Week 2
Sunday, June 26th: Keynote lecture by Darcy Peterka / Nick Steinmetz / Botond Roska Afternoon: Rig rotation
Monday, June 27th: Keynote Lecture by Marius Pachitariu / Carsen Stringer Afternoon: Miniprojects
Tuesday, June 28th: Keynote lecture by Karel Svoboda / Ana João Rodrigues Afternoon: Miniprojects
Wednesday, June 29th: Keynote lecture by Isaac Bianco Afternoon: Miniprojects
Thursday, June 30th: Keynote lecture by Michael Brecht / Marta Moita Afternoon: Miniprojects
Friday, July 1st: Keynote lecture by Megan Carey Afternoon: Miniprojects
Saturday July 2nd: Keynote lecture by Francois St. Pierre / Michael Hausser / Afternoon: Free/Social
Week 3
Sunday July 3rd: Keynote lecture by Emily Dennis / Claire Wyart / Valentina Emiliani / Afternoon: Miniprojects
Monday, July 4th: Keynote lecture by Ofer Yizhar/ Afternoon: Miniprojects
Tuesday, July 5th: Keynote lecture by Michael Orger / Afternoon: Miniprojects
Wednesday, July 6th: Keynote lecture by Ken Harris / Afternoon: Miniprojects
Thursday, July 7th: Keynote lecture by Mackenzie Mathis Afternoon: Miniprojects
Friday, July 8th: Miniproject presentation
Saturday, July 9th: Departure
Topics & Techniques
The course combines a lecture series featuring top speakers from around the world with a practical “hands-on” introduction to the latest methods for probing neural circuits, using drosophila, zebrafish, and (transgenic) mice. The course will focus on anatomy and connectivity, recording and manipulation, and the relation between circuits and behavior. During the course, each student will carry out a ‘mini-project’, executed under the guidance and supervision of experienced researchers and teaching assistants. Techniques used during the course include:
Zebrafish
- optogenetic manipulation using digital holography;
- behavior & population calcium imaging using 2-photon microscopy;
Mice
- In vivo 2-photon and 3-photon imaging;
- all-optical interrogation (simultaneous 2-photon optogenetics and 2-photon imaging);
- miniscope imaging;
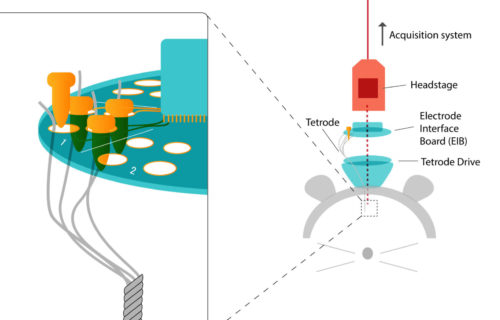
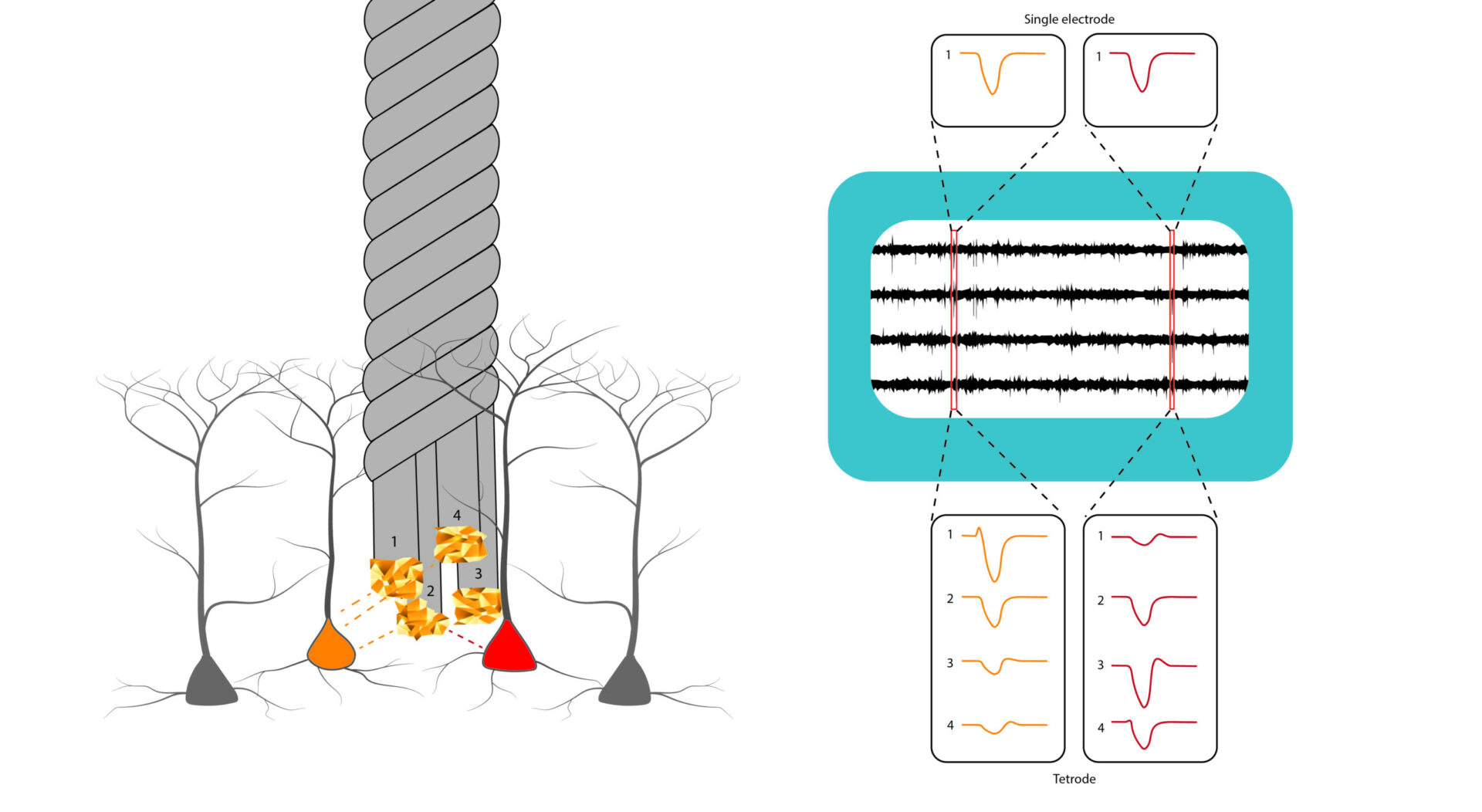
- extracellular recordings of neural population activity using Neuropixels probes in head-fixed and freely behaving animals;
- intracellular electrophysiological recordings using whole-cell patch-clamp;
- viral tracing, histology preparation, expansion microscopy, and fluorescence imaging techniques.
For more information on the course programme, you can visit the past course website.

Champalimaud Centre for the Unknown, Portugal
The Champalimaud Foundation is a private, non-profit organization, established in 2005 and dedicated to research excellence in biomedical science. Completed in 2010, the Champalimaud Centre for the Unknown is a state-of-the-art centre that houses the Champalimaud Clinical Centre and the Champalimaud Research, with its three parallel programs – the Champalimaud Neuroscience Programme, the Physiology and Cancer Programme, and the Experimental Clinical Research Programme.
Initially focused on a system and circuit approach to brain function and behavior, the Centre expanded to incorporate molecular and cell biological expertise. The Centre comprises 26 research groups (circa 400 researchers) leading independent curiosity-based research.
Facilities
The Centre provides Facilities dedicated for Training, some in their entirety, for use by the CAJAL Advanced Neuroscience Training Programme. These include the Teaching Laboratory, a fully equipped open lab space for 20-30 students that can be dynamically reconfigured to support a full range of neuroscience courses. It also overlooks, via floor to ceiling windows, a tropical garden and the river. The experimental spaces include: Imaging Lab: A dark-room containing a full size optical table is used for advanced imaging setups (two-photon microscopy, SPIM, etc.) and custom (course-designed) optical systems.
Registration
Fee : 3.500 € (includes tuition fee, accommodation and meals)
Application closed on 24th January 2022
The CAJAL programme offers 4 stipends per course (waived registration fee, not including travel expenses). Please apply through the course online application form. In order to identify candidates in real need of a stipend, any grant applicant is encouraged to first request funds from their lab, institution or government.
Kindly note that if you benefited from a Cajal stipend in the past, you are no longer eligible to receive this kind of funding. However other types of funding (such as partial travel grants from sponsors) might be made available after the participants selection pro- cess, depending on the course.
Sponsors
At Scientifica, we employ experience, collaboration, and superior design to empower you to discover the brain’s secrets and overcome neurological diseases. Our equipment is optimised for electrophysiology, multiphoton imaging and optogenetics studies.
Our highly qualified team provides first-class service and support, and our resources centre is packed with invaluable and educational content. Get in touch to see how we can help you achieve your research goals.