Course overview
Cancer neuroscience represents an interdisciplinary field at the forefront of cancer research, seamlessly combining insights from neurobiology and oncology. The intricate interplay between the nervous system and cancer has yielded profound insights into the mechanisms of tumor progression and the tumor microenvironment. The latter, comprising neuro-glial, immune, and endothelial networks, has been identified as a critical determinant of cancer growth and metastasis.
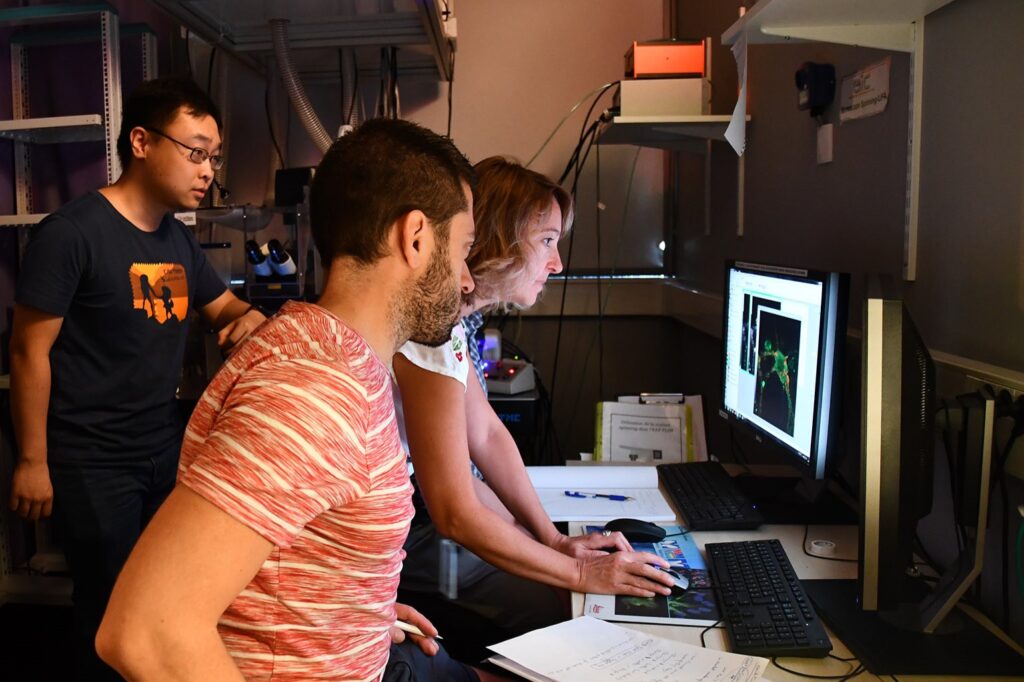
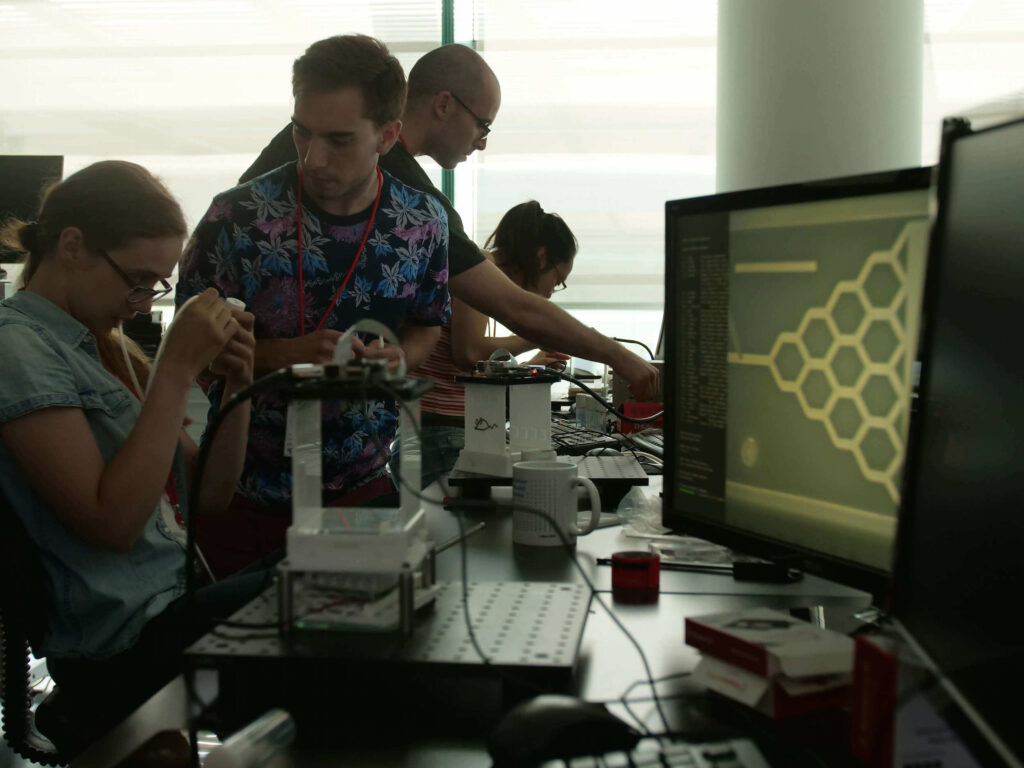
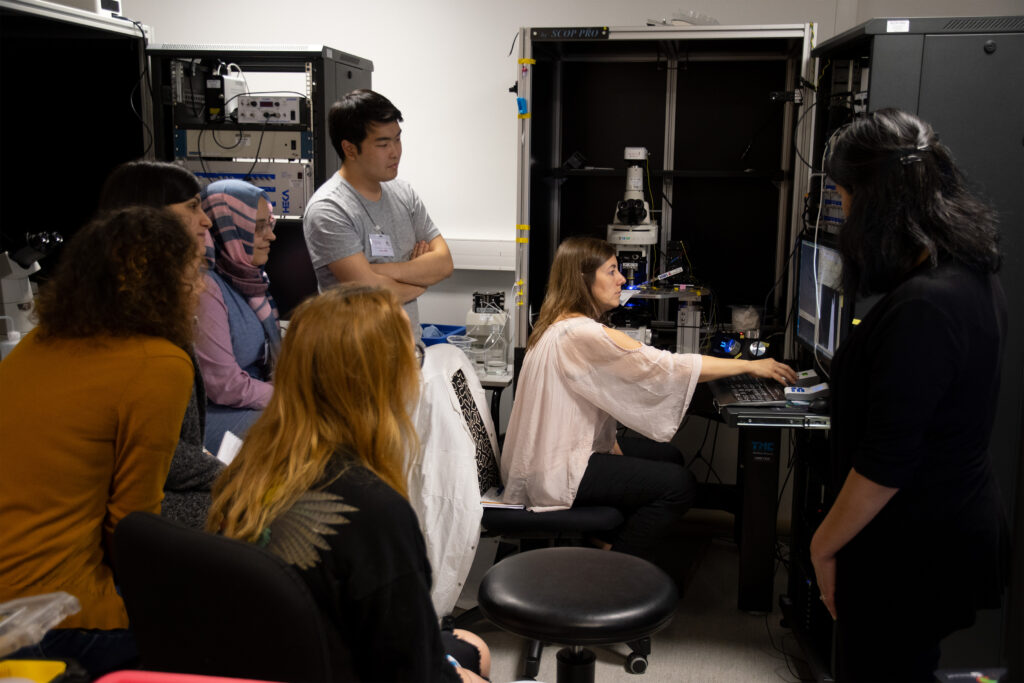
Cutting-edge technologies inspired by the neuroscience field are transforming our ability to study brain cancers from a neural perspective. This course will enable students to integrate theoretical and methodological concepts in neuro-oncology, combining modern neuroscience approaches with hands-on experience in state-of-the-art methodologies. The course provides an integrated understanding of the crosstalk between neuro-oncology and neuroscience.
Course Directors
Luxembourg Institute of Health, Luxembourg
Bordeaux Institute of Oncology – INSERM, France
German Cancer Research Center, Germany
Honorary Lecture - Brain Prize Winner
Keynote Speakers
Thomas Daubon, Institut de Biochimie et Génétique Cellulaires, France
Manuel Valiente, Centro Nacional de Investigaciones Oncológicas, Spain
Simona Parrinello, UCL Cancer Institute, UK
Aurélie Tchoghandjian, Institut de Neurophysiopathologie, France
Helene Castel, Laboratoire de Différenciation et Communication Neuronale et Neuroendocrine, France
Hrvoye Miletic, University of Bergen, Norway
Vidhya Madapusi Ravi, Freiburg Institute for Advanced Studies , Germany
Leila Akkari, Netherlands Cancer Institute, Netherlands
Dieter Henrik Heiland, Department of Neurosurgery, Germany
Elena Ciaglia, Università di Salerno, Italy
Instructors
Ahmad Charanek – Bordeaux Institute of Oncology, France
Audrey Burban – Institut de Biochimie et Génétique Cellulaires, France
Aurélie Tchoghandjia – Institut de Neurophysiopathologie, France
Bastien Redon – Neurocentre Magendie, France
Beatrice Senigagliesi – Interdisciplinary Institute for NeuroScience: Bordeaux, France
Benjamin Chauvineau – NutriNeuro, France
Camille Humeau – Bordeaux Institute of Oncology, France
Chiara Bastiancich – Institut de Neurophysiopathologie, France
Claire Larrieu – Institut de Biochimie et Génétique Cellulaires, France
Clement Morgat – Institut de Neurosciences Cognitives et Intégratives d’Aquitaine, France
Clementine Bosch Bouju – NutriNeuro, France
Cloe Tessier – Bordeaux Institute of Oncology
Doriane Bomont – Institut de Biochimie et Génétique Cellulaires, France
Ekin Reyhan – Universität Heidelberg, Germany
Emanuelle Georget – Bordeaux Institute of Oncology
Kevin Boye – Paris Cardiovascular Research Center, France
Kirill Smirnov – Institut de Psychiatrie et Neurosciences de Paris, France
Lucie Brisson – Bordeaux Institute of Oncology, France
Mahsa Rezaeipour – Luxembourg Institute of Health, Luxembourg
Maialen Arrieta – Institut de Biochimie et Génétique Cellulaires, France
Maria Haykal – Institut de Biochimie et Génétique Cellulaires, France
Mathis Pinglaut – Institut de Biochimie et Génétique Cellulaires, France
Nassim Haffiane – Interdisciplinary Institute for NeuroScience: Bordeaux, France
Oceane Martin – Institut de Biochimie et Génétique Cellulaires, France
Pilar Moreno-Sanchez – Luxembourg Institute of Health, Luxembourg
Stella Soyka – Universität Heidelberg, Germany
Thomas Daubon – Institut de Biochimie et Génétique Cellulaires, France
Valentina Lopardo – Università di Salerno, Italy
Course Content and Techniques
Cultures of brain tumor cells
Primary cultures of neurons, astrocytes, and oligodendrocytes
Primary culture of enteric nervous system cells (ENS) and ex vivo analysis of ENS structure in mouse models
Tumor cell spheroid cultures
Organoids
Organotypic brain slices
Co-culture of tumor cells with brain cells
Culture of glioblastoma cells in 3D-living systems of brain
Stereotaxic surgery for tumor implantation
Tumor debulking via the biopsy punch technique
Live imaging of cell-cell interactions
Confocal microscopy
Spinning-disk microscopy
Super-resolution microscopy
Two-photon microscopy
Multiphoton longitudinal imaging of tumor progression
DREADD manipulation and calcium imaging in mice during reward-based navigation.
Neuronal calcium activity using a head-mounted miniature microscope on animals performing behavioral tasks
Magnetic Resonance Imaging
Radiolabeling with radiometals and characterization of radio pharmaceuticals
SPECT/CT imaging and post-mortem biodistribution
PBMC isolation from human blood
Isolation of murine tumor-infiltrating immune cells
Characterization of immune cells by flow cytometry
High resolution respirometry using Resipher system and Oroboros Oxygraph-2k
Isolation of proteins and exosomes.
PamGene Technology
Methods to induce Blood-brain barrier opening in brain tumor models
Electrophysiology in multielectrode array
Electrophysiological patch-clamp recordings
Introduction to signal preprocessing, spike detection, and activity mapping
Image analysis using Imaris software
Analysis of cell motility with ImageJ
Analysis of spatial transcriptomics datasets using Seurat, Squidpy, R and Python
Analysis of 10x Genomics Visium datasets from human glioblastoma samples
Exploration of 10x Genomics Xenium imaging-based spatial data

Bordeaux School of Neuroscience, France
The Bordeaux School of Neuroscience is part of Bordeaux Neurocampus, the Neuroscience Department of the University of Bordeaux. Created in 2015, it is currently directed by David Perrais. Throughout the year, renowned scientists, promising young researchers and many students from any geographical horizon come to the School.
The school works on this principle: training in neuroscience research through experimental practice, within the framework of a real research laboratory.
Facilities
Their dedicated laboratory (500m2), available for about 20 trainees, is equipped with a wet lab, an in vitro and in vivo electrophysiology room, IT facilities, a standard cellular imaging room, an animal facility equipped for behavior studies and surgery and catering/meeting spaces. They also have access to high-level core facilities within the University of Bordeaux. They offer their services to international training teams who wish to organize courses in all fields of neuroscience thanks to a dedicated staff for the full logistics (travels, accommodation, on-site catering, social events) and administration and 2 scientific managers in support of the experimentation.

Registration
Fee : 4 500 € (includes tuition fee, accommodation and meals)
Applications are closed.
The CAJAL programme offers 4 stipends per course (waived registration fee, not including travel expenses). Please apply through the course online application form. In order to identify candidates in real need of a stipend, any grant applicant is encouraged to first request funds from their lab, institution or government.
Kindly note that if you benefited from a Cajal stipend in the past, you are no longer eligible to receive this kind of funding. However other types of funding (such as partial travel grants from sponsors) might be made available after the participants selection pro- cess, depending on the course.